|
RE sites for the MCR design were determined by the trial-anderror Possible to clone PCR products directly into linearized T-vector. Placed in such a way that a digestion of the modified vector withĮndonuclease XcmI could give rise to 3’-Ts overhanging on DNAĭuplexes complementary to adenines of the PCR products. Oligonucleotide as a polylinker with two XcmI recognition The main idea of our experiment was preparing of double-stranded Materials and Method: Construction of Multi Cloning Region for the GeneClip vector: The insert from one plasmid to the others. That can be used both for cloning PCR products and transferring To overcome these difficulties through design of universal MCR Variants or confirmations with a different object or insert. Very handy when the experiment requires a number of repeats and 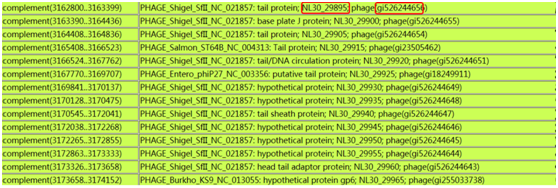
Molecular biology procedure, it claims a lot of time, and is not To select a suitable site for subcloning the insert from a subsidiary However sets of RS are often limited and it is difficult T-ends of such T-vectors are surroundedīy MCRs containing RE sites that can be used for the inserts There is a variety of T-vector systems designed for efficientĬloning of PCR products. With dATP and a nonproofreading DNA polymerase. Blunt-ended productsĭerived from PCR with proofreading Pfu and Tli polymerases canīe modified with 3’-overhangs by incubating the PCR fragment PCR products allow to clone them directly into linearized T-vectorsĪvoiding restriction and ligation. Overhanging on DNA duplexes complementary to adenines of the Proofreading activity but can add a single base extention ofĭeoxyadenosine to the 3´-ends of amplified products. ThermostableĭNA polymerases such as Taq, Tth, Tfl are most often used toĪmplify the DNA fragments. Modified PCR applications appeared to perform a wide range ofĭifferent DNA polymerases can be used to optimize DNAĪmplification depending on the aim of experiment. Since polymerase chain reaction was invented new Magnitude, generating the multitude of copies of a particular DNA The polymerase chain reaction (PCR) method enables one toĪmplify a single DNA (or RNA) segment across several orders of Keywords: PCR Products Cloning, T-vector, subcloning, Multiple Cloning Region (MCR), Restriction Sites (RS) So, everybody can now construct MCRs for their own vectors using this manual. Here, we elaborated design of the MCR for a GeneClip vector that can be used both for cloning PCR products and transferring the Method is that a lot of MCRs contain a limited number of Restriction Sites (RS), which often makes the process of subcloning arduous. The main problem of such the trial-and-error Most of them containĭifferent Multiple Cloning Regions (MCR)s depending on specificity of the experiments. It is the best environment for molecular biology.There exist a lot of vectors for a wide range of experiments, and new modified plasmids appear every year. It is compatible with both x86 and 圆4 architecture. SnapGene Portable 3.2.1 Free Download for WindowsĬlicking the below button will start downloader the standalone portable version of SnapGene 3.2.1 for Windows. Compatible with Windows 10/8/7/Vista/XP.Take a look at the technical details of SnapGene before downloading it. 
Technical Details of Portable SnapGene 3.2.1 Export sequence maps or specific section. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
